Scaling from molecules to ecosystems—Cladophora epiphytes as a model microbiome
Cladophora is the dominant filamentous green algae in the Eel River (and in temperate waters worldwide) (Fig. 1). It has a rough “skin” of cellulose-rich cell walls, so becomes colonized by many microorganisms and small animals called epiphytes (Fig. 2).
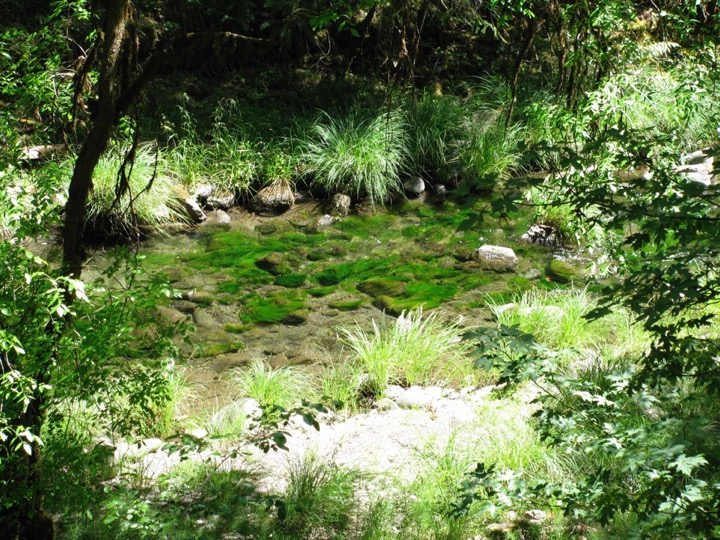 Fig. 1
Fig. 1
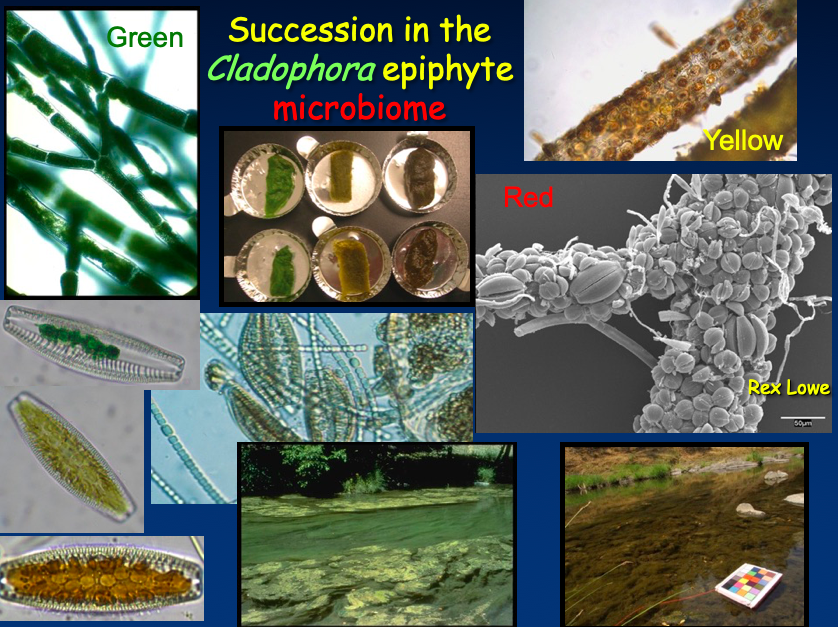 Fig. 2
Fig. 2
In an NSF “Rules of Life” project led by Jane Marks (a Northern Arizona University professor and an Eel River alumna), we are examining the microbiome hosted on Cladophora in the Eel River. Using molecular techniques like qSIP (quantitative stable isotope probing) and nanoSIMS (Nanoscale secondary ion mass spectrometry), Jane’s team can track growth rates of members of Cladophora’s microbiome under natural or manipulated light, flow velocity, temperature, or nutrient levels. We can also follow elements through the microbiome food web, as members compete with, facilitate, or eat each other. For example, Jane and colleagues at the Lawrence Livermore National Lab have used nanoSIMS to follow atmospheric nitrogen fixed by cyanobacterial endosymbionts of the diatom Epithemia, tracing N from the air into the endosymbiont, into the host diatom’s cytoplasm, then outside that cell into the microbiome matrix, and finally into neighboring diatoms that don’t fix nitrogen (Fig. 3, lower left).
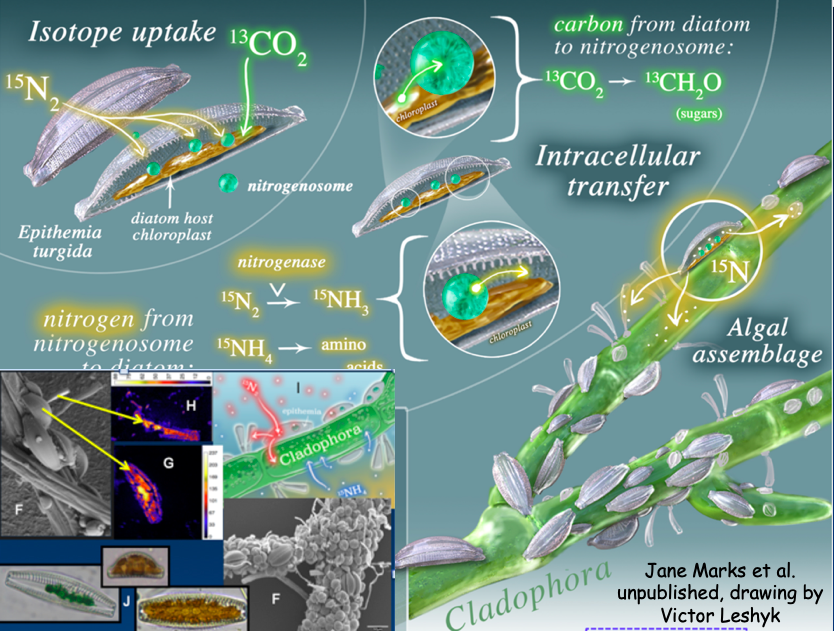 Fig. 3
Fig. 3
Studying the ecology of microbiomes has become a new frontier for biology, and rules of engagement may be easier to discover in the visible and manipulable Cladophora epiphyte microbiome than in the microbiomes of guts, mouths, or soils, where they have received more scientific attention.
|

