|
Evolutionary
Biology of the Australian Wet Tropics
Molecular evidence supports the contention that historical
contraction and expansion of the Australian Wet Tropics (AWT)
rainforest has affected species distributions and diversification,
but also provides qualitative evidence for differences among
species in the temporal and spatial scale of their responses.
We combine ecological, GIS and molecular methods to examine
the spatial history of amphibian and reptile species endemic
to the biologically diverse and intensively studied rainforests
of the Australian Wet Tropics. Our long-term aim for this
system is to understand how changes in rainforest distribution
have influenced speciation and adaptive divergence (Moritz
et al. 2000; Moritz 2002) and how these processes, along with
local extinction, recolonisation and species interactions
shape patterns of community diversity (Williams et al. 1996,
Williams 1997, Williams & Pearson 1997, Williams &
Hero 2001, Schneider & Williams in press). Our research
to date has documented patterns of community diversity and
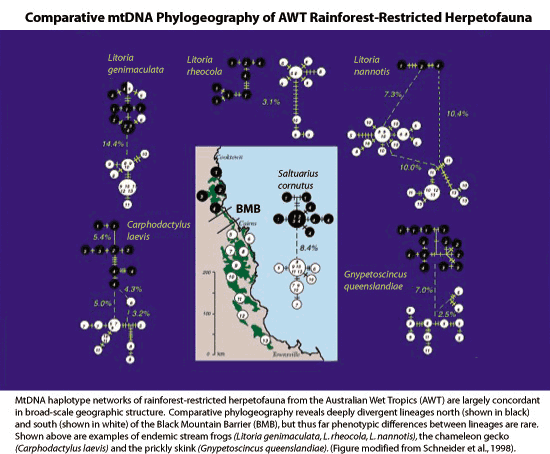
comparative phylogeography. Phylogeographic studies of rainforest-restricted
fauna in the AWT frequently reveal spatially congruent clade
distributions and secondary contact zones (Joseph et al. 1995;
Schneider et al. 1998), offering strong qualitative agreement
with the locations of historical refugia and predicted areas
of range expansion events in climatic paleomodelling (Hugall
et al. 2002). However, levels of observed (net) sequence divergence
across shared biogeographic barriers (eg. the Black Mountain
Barrier) vary widely even within a genus and several species
show additional genetic breaks across large river drainages
(eg. Bloomfield, Tully). A major aim of our current research
is to understand how and why species vary in their responses
to a common history of rainforest fluctuation. It is clear
that more precise and spatially explicit estimates of population
history across co-distributed species are needed. Further,
we need to integrate the molecular analyses with information
on relevant ecological attributes of each species if we are
to predict and understand their responses to spatial models
of past and future habitat change.
Our current approach combines multi-locus estimates of population
history with spatially explicit models of habitat distributions
under varying climates and with field-derived estimates of
habitat specialization and climate “niche” breadth.
Novel aspects of the research include the following: 1) We
will apply multilocus coalescent methods
to infer population divergence, size(s) and migration rates
across multiple codistributed species in a system with a well-known
history of habitat fluctuations. Multilocus approaches have
not previously been applied to comparative biogeography on
this scale. 2) We will develop explicit spatial models of
historical distributions of species by combining the molecular
evidence with paleodistribution models that incorporate information
on climate-driven changes in habitat distributions, conditioned
on observed environmental niche breadth and dispersal limitation.
3) By combining the models of individual species, we will
generate spatial models of assemblage structure through the
climatic fluctuations of the late Quaternary. 4) We will use
our knowledge of sensitivity of species to historical climate
change to improve predictions about the impacts that future
climate change on potential ranges of these species.
|
 |